Functional Neuroimaging and Biophotonics Lab
Projects
Wide Field Optical Imaging of Brain Activity
Goal: Map functional brain organization in the mouse
Functionally-related brain regions exhibit correlated neural and hemodynamic activity even in the absence of imposed tasks or overt motor movements (i.e. under “resting-state” conditions). This correlated spontaneous activity defines widely-distributed topographies known as resting state networks or resting-state functional connectivity (FC) patterns. Importantly, patterns of FC are indicative of bran integrity; changes in FC have been reported in a variety of neuro-psychiatric disorders including Alzheimer’s disease, depression, schizophrenia, Parkinson’s disease and stroke. Because basic functional connectional topology is conserved across mice, rats, primates, and humans, bridge measurements can be made across animal models to inform and enrich findings in humans.
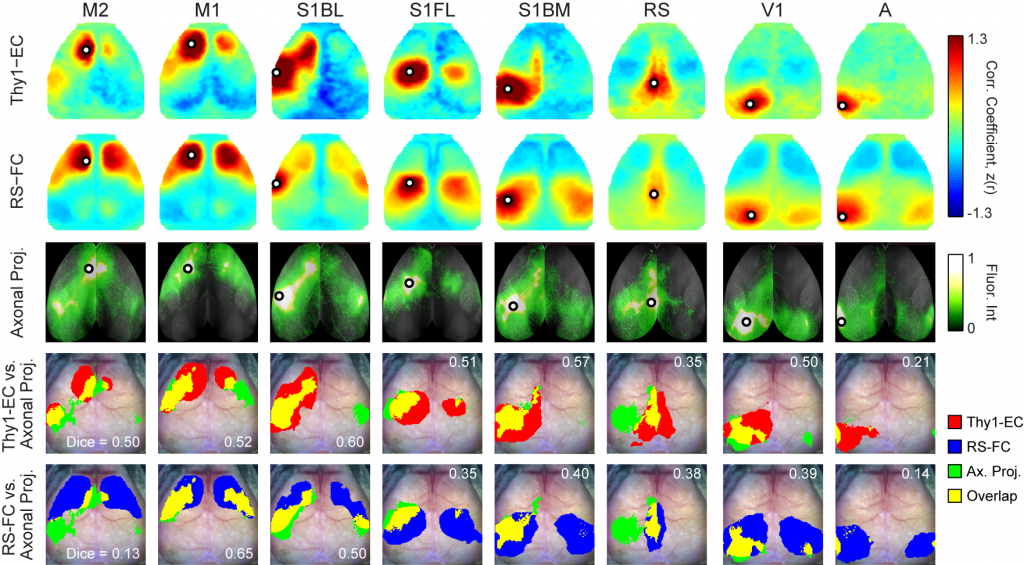
Optogenetic Mapping of Brain Circuitry
Goal: Determine cell-specific patterns of cortical connectivity in mice
Originally conceptualized, the notion of brain connectomics referred to axonal connectivity, i.e., white matter tracts, classically studied using histological methods. More recently, the concept of connectomics has expanded to also encompass resting state functional connectivity (FC) described above, as well as effective connectivity (EC). EC is distinct from, but related to, both anatomical and FC. EC measures the influence (direct or indirect) that one brain region exerts on another. The crucial distinction between FC vs. EC is that FC characterizes shared spontaneous (ongoing, intrinsic) signals. By definition, pairwise FC is symmetric and uninformative regarding the directionality of propagated signals. In contrast, EC reports how activity in an identified part of the brain affects other regions. Thus, measures of EC are not necessarily symmetric, and instead capable of revealing aspects of functional connectivity obscured by network-level synchronization.
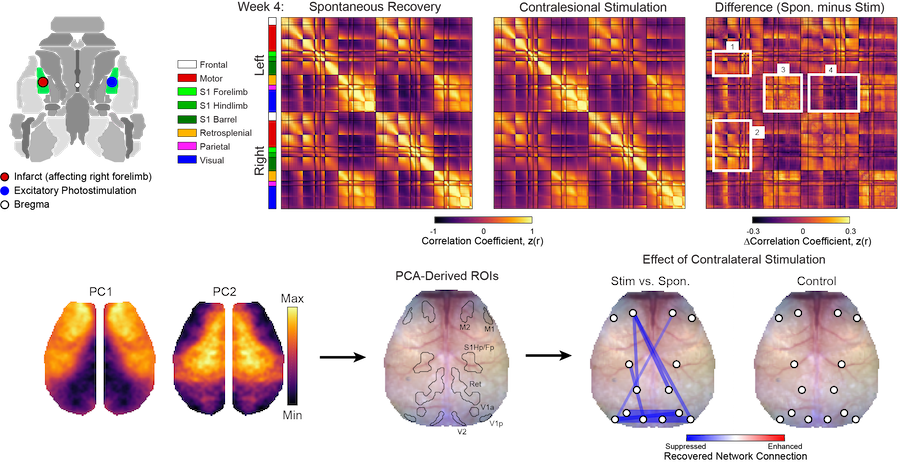
Manipulating Functional Recovery After Stroke
Goal: Understand endogenous mechanisms of brain repair after focal ischemia
Stroke causes direct structural damage to local circuits and indirect functional damage to global networks that can result in behavioral deficits spanning multiple domains. Neuroplasticity after stroke involves molecular changes within perilesional tissue that can be influenced by distant regions spared from injury. We are using a variety of manipulations to understand how physical therapy (e.g., multisensory stimulation), neural activity (e.g., via optogenetic targeting) and molecular mechanisms of repair (e.g., axonal sprouting cues) influence local circuit repair (e.g., cortical remapping and restoration of monosynaptic connections) and global network reintegration (recovery of FC) after stroke in mice.
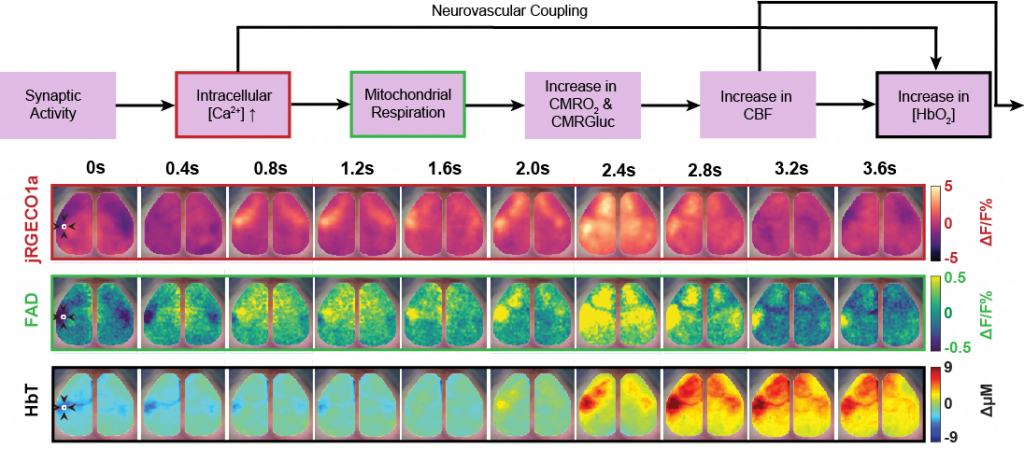
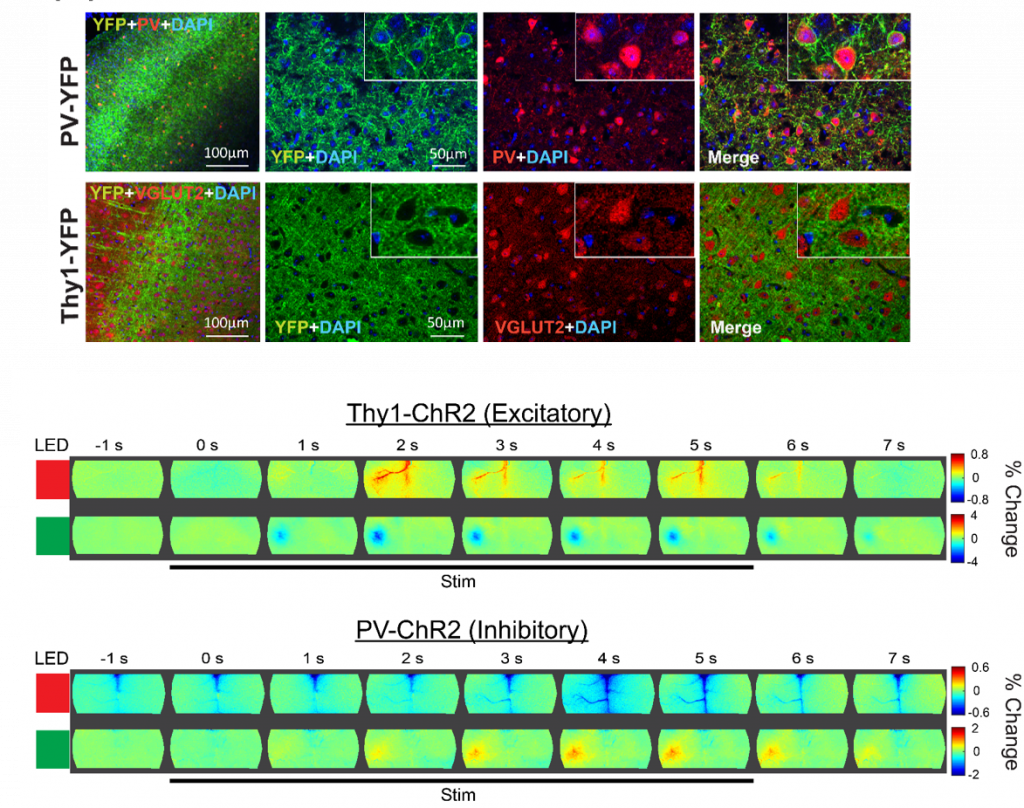
Cellular Contributions to Neurovascular Coupling
Goal: Understand how activity in different cell population contributes to local and distant changes in cerebral blood flow.
Neurovascular coupling (NVC) is the process through which changes in local neural activity are coupled to changes in cerebral blood flow. The magnitude and location of these hemodynamic changes are tightly linked to changes in neural activity through a series of coordinated events involving neurons, glia, and vascular cells. Understanding how underlying electrophysiological and/or metabolic activity relates to the local hemodynamic response is essential for interpreting blood-based mapping signals. An important link connecting neural activity and blood flow regulation is the ability of certain cell populations to mediate the diameter of local blood vessels through vasoactive messengers. Across several new projects, we are combining optogenetic targeting with optical intrinsic signal imaging and wide-field calcium fluorescence imaging to study how activity in different cell population contributes to NVC.
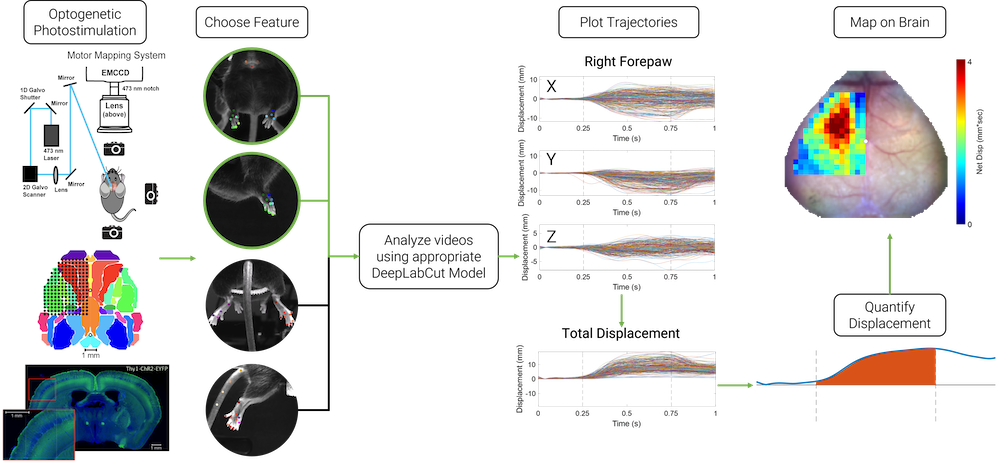
Light-Based Motor Mapping Using Deep Learning
Goal: Map fine motor movements of multiple limbs in 3D
Motor mapping allows for determining the macroscopic organization of motor circuits and corresponding motor movement representations on the cortex. Methods such as intracortical microstimulation can produce reliable motor maps, but the technique is time consuming and invasive, making it challenging to perform in longitudinal studies. Optogenetic motor mapping, alternatively known as light-based motor mapping, provides a rapid and minimally invasive technique for examining cortical motor movement representations compared to intracortical microstimulation, but has seen limited use in tracking behaviorally-relevant movement trajectories of multiple limbs. We use deep learning conjunction with multiple feature tracking in mice to localize the movements of multiple limbs concurrently. This method allows for quick, minimally invasive, longitudinal mapping, and can be used for monitoring functional recovery from brain injury.