Neuroinformatics Research Group
Projects
XNAT
Open-Source Software Platform
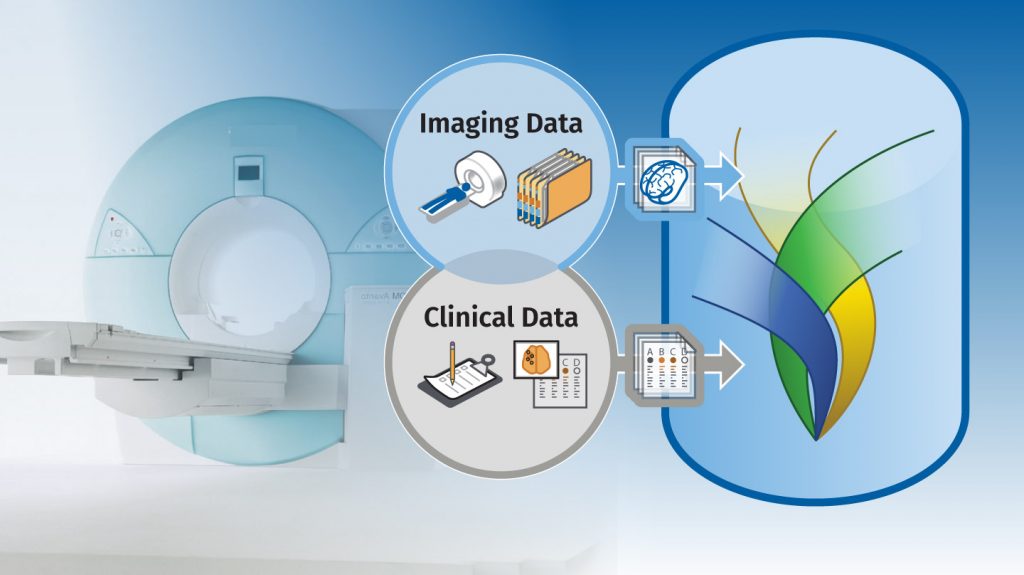
XNAT is an open-source software platform developed to support imaging informatics research. The platform offers permission-controlled storage of imaging and clinical assessment data, as well as support for containerized data processing to support reproducible science. XNAT has been used in hundreds of research projects and clinical trials at institutions around the world.
Human Connectome Project
Mapping the Brain Across the Human Lifespan
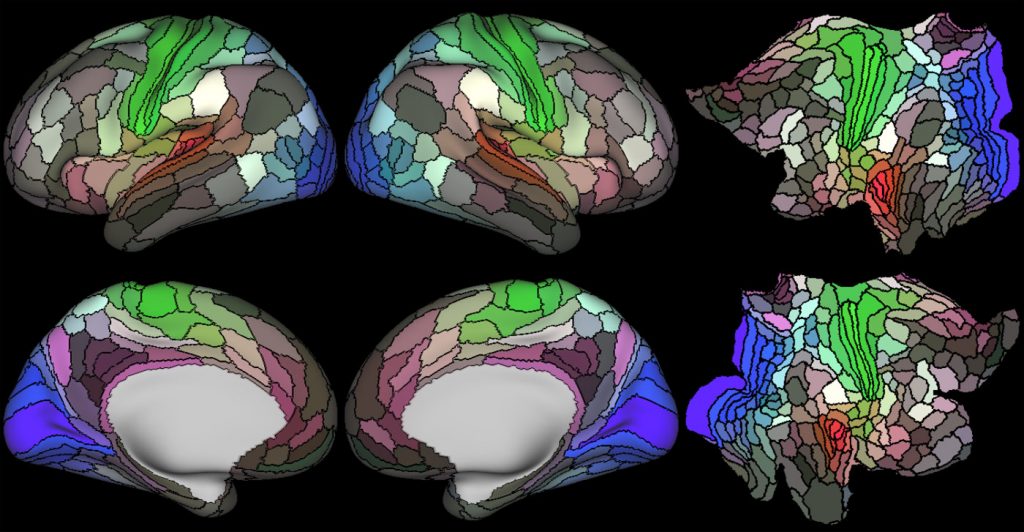
The Human Connectome Project (HCP) encompasses over 20 funded studies to map the brain across the human lifespan and in a range of neurological and psychiatric disorders. The initial HCP Young Adult study imaged more than 1100 young using a cutting edge scanner and set of structural and functional sequences. The resulting data set was shared openly with the research community through the XNAT-based ConnectomeDB platform. Since the end of the original HCP grant, the HCP operations in the NRG has been supported by the Connectome Coordinating Facility (CCF), enabling 19 other disease and aging related studies. Ongoing study of the aging brain continues with the Aging Adult Brain Connectome (AABC)
OASIS
Open Access Series of Imaging Studies
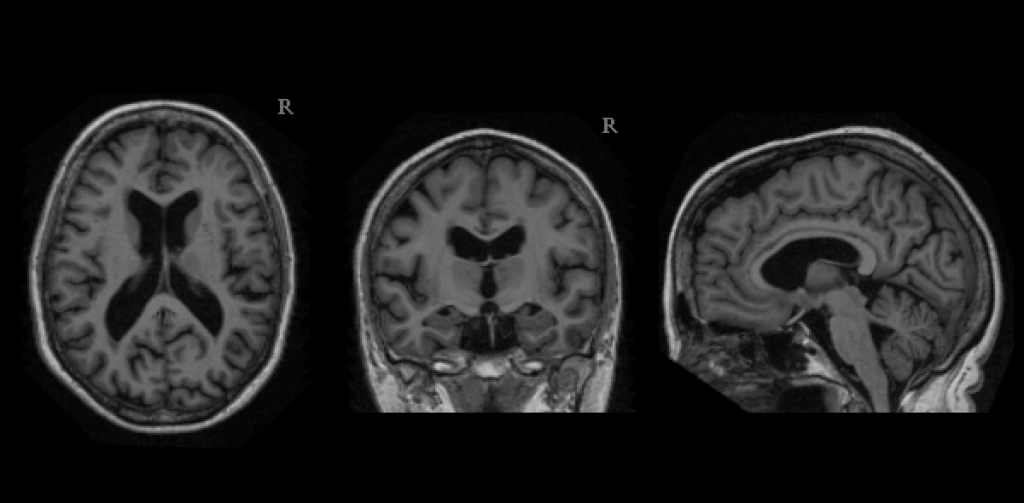
OASIS is a project aimed at making neuroimaging data sets of the brain freely available to the scientific community. By compiling and freely distributing neuroimaging data sets, we hope to facilitate future discoveries in basic and clinical neuroscience.
MIR Research Imaging Repository
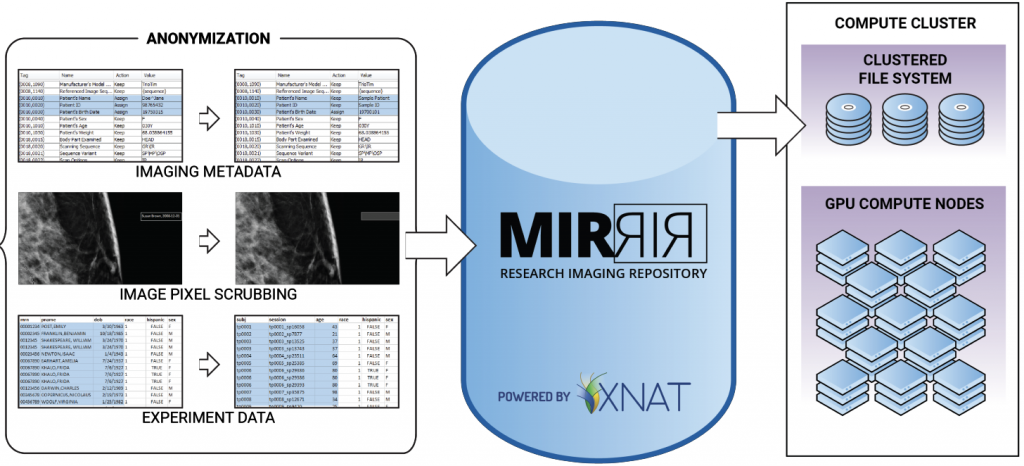
The MIR Research Image Repository (MIRRIR) is a datastore consisting of radiological images and reports used by Washington University investigators to conduct translational research. This includes development for computational algorithms for disease prediction, diagnosis and treatment guidance. MIRRIR is designed to hold all radiology exams obtained in BJC HealthCare hospitals and the associated reports and observations generated by MIR radiologists.
Translational Imaging Portal
Subhead
The goal of translational medicine is to speed the application of analytic methods developed in the lab into clinical practice. The Translational Imaging Portal (TIP) is the result of a collaboration between the CIRC and radiologists at Barnes-Jewish Hospital. What started as a pilot program and proof-of-concept in 2015 is now a continuing part of surgical planning at BJC HealthCare.
Integrative Imaging Informatics for Cancer Research Center
The Integrative Imaging Informatics for Cancer Research Center (I3CR) focuses on expanding the open source XNAT informatics platform to better support computational workflows in cancer imaging. I3CR has also developed knowledge management tools to better track data processing and analysis, including tools for orchestrating and tracking container-based computing pipelines.